Copy number calling pipeline¶
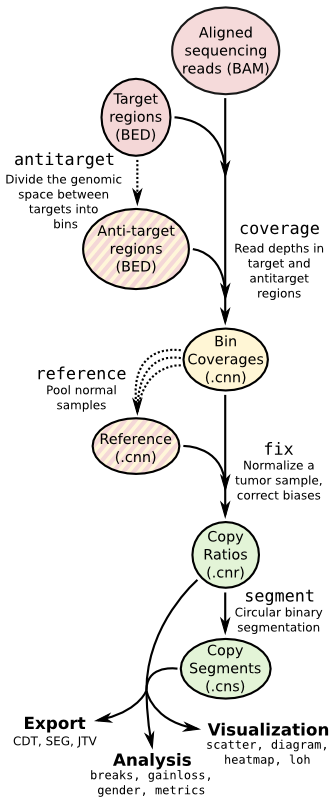
Each operation is invoked as a sub-command of the main script, cnvkit.py.
A listing of all sub-commands can be obtained with cnvkit --help or -h,
and the usage information for each sub-command can be shown with the --help
or -h option after each sub-command name:
cnvkit.py -h
cnvkit.py target -h
A sensible output file name is normally chosen if it isn’t specified, except in
the case of the text reporting commands, which print to standard output by
default, and the matplotlib-based plotting commands (not diagram), which
will display the plots interactively on the screen by default.
batch¶
Run the CNVkit pipeline on one or more BAM files:
# From baits and tumor/normal BAMs
cnvkit.py batch *Tumor.bam --normal *Normal.bam \
--targets my_baits.bed --split --annotate refFlat.txt \
--fasta hg19.fasta --access data/access-5kb-mappable.hg19.bed \
--output-reference my_reference.cnn --output-dir results/ \
--diagram --scatter
# Reusing a reference for additional samples
cnvkit.py batch *Tumor.bam -r Reference.cnn -d results/
# Reusing targets and antitargets to build a new reference, but no analysis
cnvkit.py batch -n *Normal.bam --output-reference new_reference.cnn \
-t my_targets.bed -a my_antitargets.bed --male-reference \
-f hg19.fasta -g data/access-5kb-mappable.hg19.bed
With the -p option, process each of the BAM files in parallel, as separate
subprocesses. The status messages logged to the console will be somewhat
disorderly, but the pipeline will take advantage of multiple CPU cores to
complete sooner.
cnvkit.py batch *.bam -r my_reference.cnn -p 8
The pipeline executed by the batch command is equivalent to:
cnvkit.py target baits.bed [--split --annotate --short-names] -o my_targets.bed
cnvkit.py antitarget my_targets.bed [--access] -o my_antitargets.bed
# For each sample...
cnvkit.py coverage Sample.bam my_targets.bed -o Sample.targetcoverage.cnn
cnvkit.py coverage Sample.bam my_antitargets.bed -o Sample.antitargetcoverage.cnn
# With all normal samples...
cnvkit.py reference *Normal.bam -t my_targets.bed -a my_antitargets.bed \
[--fasta hg19.fa --male-reference] -o my_reference.cnn
# For each tumor sample...
cnvkit.py fix Sample.targetcoverage.cnn Sample.antitargetcoverage.cnn my_reference.cnn -o Sample.cnr
cnvkit.py segment Sample.cnr -o Sample.cns
# Optionally, with --scatter and --diagram
cnvkit.py scatter Sample.cnr -s Sample.cns -o Sample-scatter.pdf
cnvkit.py diagram Sample.cnr -s Sample.cns [--male-reference] -o Sample-diagram.pdf
See the rest of the commands below to learn about each of these steps and other functionality in CNVkit.
target¶
Prepare a BED file of baited regions for use with CNVkit.
cnvkit.py target my_baits.bed --annotate refFlat.txt --split -o my_targets.bed
The BED file should be the baited genomic regions for your target capture kit,
as provided by your vendor. Since these regions (usually exons) may be of
unequal size, the --split option divides the larger regions so that the
average bin size after dividing is close to the size specified by
--average-size. If any of these three (--split, --annotate, or
--short-names) flags are used, a new target BED file will be created;
otherwise, the provided target BED file will be used as-is.
Bin size and resolution¶
If you need higher resolution, you can select a smaller average size for your target and antitarget bins.
Exons in the human genome have an average size of about 200bp. The target bin size default of 267 is chosen so that splitting larger exons will produce bins with a minimum size of 200. Since bins that contain fewer reads result in a noisier copy number signal, this approach ensures the “noisiness” of the bins produced by splitting larger exons will be no worse than average.
Setting the average size of target bins to 100bp, for example, will yield about twice as many target bins, which might result in higher-resolution segmentation. However, the number of reads counted in each bin will be reduced by about half, increasing the variance or “noise” in bin-level coverages. An excess of noisy bins can make visualization difficult, and since the noise may not be Gaussian, especially in the presence of many bins with zero reads, the CBS algorithm could produce less accurate segmentation results on low-coverage samples. In practice we see good results with an average of 200-300 reads per bin; we therefore recommend an overall on-target sequencing coverage depth of at least 200x to 300x with a read length of 100 to justify reducing the average target bin size to 100bp.
Adding gene names¶
In case the vendor BED file does not label each region with a corresponding gene
name, the --annotate option can add or replace these labels.
Gene annotation databases, e.g. RefSeq or Ensembl, are available in “flat”
format from UCSC (e.g. refFlat.txt for hg19).
In other cases the region labels are a combination of human-readable gene names
and database accession codes, separated by commas (e.g.
“ref|BRAF,mRNA|AB529216,ens|ENST00000496384”). The --short-names option
splits these accessions on commas, then chooses the single accession that covers
in the maximum number of consecutive regions that share that accession, and
applies it as the new label for those regions. (You may find it simpler to just
apply the refFlat annotations.)
access¶
Calculate the sequence-accessible coordinates in chromosomes from the given reference genome, output as a BED file.
cnvkit.py access hg19.fa -x excludes.bed -o access-hg19.bed
Many fully sequenced genomes, including the human genome, contain large regions of DNA that are inaccessable to sequencing. (These are mainly the centromeres, telomeres, and highly repetitive regions.) In the FASTA reference genome sequence these regions are filled in with large stretches of “N” characters. These regions cannot be mapped by resequencing, so we will want to avoid them when calculating the antitarget bin locations (for example).
The access command computes the locations of the accessible sequence regions
for a given reference genome based on these masked-out sequences, treating long
spans of ‘N’ characters as the inaccessible regions and outputting the
coordinates of the regions between them.
Other known unmappable or poorly sequenced regions can be specified for
exclusion with the -x option.
This option can be used more than once to exclude several BED files listing
different sets of regions.
For example, “excludable” regions of poor mappability have been precalculated by
others and are available from the UCSC FTP Server (see here for hg19).
If there are many small excluded/inaccessible regions in the genome, then small,
less-reliable antitarget bins would be squeezed into the remaining accessible
regions. The -s option tells the script to ignore short regions that would
otherwise be excluded as inaccessible, allowing larger antitarget bins to
overlap them.
An “access” file precomputed for the UCSC reference human genome build hg19,
with some know low-mappability regions excluded, is included in the CNVkit
source distribution under the data/ directory
(data/access-5kb-mappable.hg19.bed).
antitarget¶
Given a “target” BED file that lists the chromosomal coordinates of the tiled regions used for targeted resequencing, derive a BED file off-target/”antitarget”/”background” regions.
cnvkit.py antitarget my_targets.bed -g data/access-5kb-mappable.hg19.bed -o my_antitargets.bed
Certain genomic regions cannot be mapped by short-read resequencing (see
access); we can avoid them when calculating the antitarget locations by
passing the locations of the accessible sequence regions with the -g or
--access option. CNVkit will then compute “antitarget” bins only within the
accessible genomic regions specified in the “access” file.
CNVkit uses a cautious default off-target bin size that, in our experience, will typically include more reads than the average on-target bin. However, we encourage the user to examine the coverage statistics reported by CNVkit and specify a properly calculated off-target bin size for their samples in order to maximize copy number information.
Off-target bin size¶
An appropriate off-target bin size can be computed as the product of the average target region size and the fold-enrichment of sequencing reads in targeted regions, such that roughly the same number of reads are mapped to on– and off-target bins on average — roughly proportional to the level of on-target enrichment.
The preliminary coverage information can be obtained with the script CalculateHsMetrics in the Picard suite (http://picard.sourceforge.net/), or from the console output of the CNVkit coverage command when run on the target regions.
coverage¶
Calculate coverage in the given regions from BAM read depths.
With the -p option, calculates mean read depth from a pileup; otherwise, counts the number of read start positions in the interval and normalizes to the interval size.
cnvkit.py coverage Sample.bam Tiled.bed -o Sample.targetcoverage.cnn
cnvkit.py coverage Sample.bam Background.bed -o Sample.antitargetcoverage.cnn
Summary statistics of read counts and their binning are printed to standard error when CNVkit finishes calculating the coverage of each sample (through either the batch or coverage commands).
BAM file preparation¶
For best results, use an aligner such as BWA-MEM, with the option to mark secondary mappings of reads, and flag PCR duplicates with a program such as SAMBLASTER, SAMBAMBA, or the MarkDuplicates script in Picard tools, so that CNVkit will skip these reads when calculating read depth.
You will probably want to index the finished BAM file using samtools or SAMBAMBA. But if you haven’t done this beforehand, CNVkit will automatically do it for you.
Note
The BAM file must be sorted. CNVkit will check that the first few reads are sorted in positional order, and raise an error if they are not. However, CNVkit might not notice if reads later in the file are unsorted; it will just silently ignore the out-of-order reads and the coverages will be zero after that point. So be safe, and sort your BAM file properly.
Note
If you’ve prebuilt the BAM index file (.bai), make sure its timestamp is
later than the BAM file’s. CNVkit will automatically index the BAM file
if needed – that is, if the .bai file is missing, or if the timestamp
of the .bai file is older than that of the corresponding .bam file. This
is done in case the BAM file has changed after the index was initially
created. (If the index is wrong, CNVkit will not catch this, and coverages
will be mysteriously truncated to zero after a certain point.) However,
if you copy a set of BAM files and their index files (.bai) together over
a network, the smaller .bai files will typically finish downloading first,
and so their timestamp will be earlier than the corresponding BAM or FASTA
file. CNVkit will then consider the index files to be out of date and will
attempt to rebuild them. To prevent this, use the Unix command touch
to update the timestamp on the index files after all files have been
downloaded.
reference¶
Compile a copy-number reference from the given files or directory (containing normal samples). If given a reference genome (-f option), also calculate the GC content of each region.
cnvkit.py reference -o Reference.cnn -f ucsc.hg19.fa *targetcoverage.cnn
The reference can be constructed from zero, one or multiple control samples. A reference should be constructed specifically for each target capture panel (i.e. set of baits) and, ideally, match the type of sample (e.g. FFPE-extracted or fresh DNA) and library preparation protocol or kit used.
Paired or pooled normals¶
To analyze a cohort sequenced on a single platform, we recommend combining all normal samples into a pooled reference, even if matched tumor-normal pairs were sequenced – our benchmarking showed that a pooled reference performed slightly better than constructing a separate reference for each matched tumor-normal pair. Furthermore, even matched normals from a cohort sequenced together can exhibit distinctly different copy number biases (see Plagnol et al. 2012 and Backenroth et al. 2014); reusing a pooled reference across the cohort provides some consistency to help diagnose such issues.
Notes on sample selection:
- You can use
cnvkit.py metrics *.cnr -s *.cnsto see if any samples are especially noisy. See the metrics command. - CNVkit will usually call larger CNAs reliably down to about 10x on-target coverage, but there will tend to be more spurious segments, and smaller-scale or subclonal CNAs can be hard to infer below that point. This is well below the minimum coverage thresholds typically used for SNV calling, especially for targeted sequencing of tumor samples that may have significant normal-cell contamination and subclonal tumor-cell populations. So, a normal sample that passes your other QC checks will probably be OK to use in building a CNVkit reference – assuming it was sequenced on the same platform as the other samples you’re calling.
If normal samples are not available, it will sometimes be acceptable to build the
reference from a collection of tumor samples. You can use the scatter command
on the raw .cnn coverage files to help choose samples with relatively
minimal and non-recurrent CNVs for use in the reference.
With no control samples¶
Alternatively, you can create a “flat” reference of neutral copy number (i.e. log2 0.0) for each probe from the target and antitarget interval files. This still computes the GC content of each region if the reference genome is given.
cnvkit.py reference -o FlatReference.cnn -f ucsc.hg19.fa -t Tiled.bed -a Background.bed
Possible uses for a flat reference include:
- Extract copy number information from one or a small number of tumor samples when no suitable reference or set of normal samples is available. The copy number calls will not be quite as accurate, but large-scale CNVs should still be visible.
- Create a “dummy” reference to use as input to the
batchcommand to process a set of normal samples. Then, create a “real” reference from the resulting*.targetcoverage.cnnand*.antitargetcoverage.cnnfiles, and re-runbatchon a set of tumor samples using this updated reference. - Evaluate whether a given paired or pooled reference is suitable for an analysis by repeating the CNVkit analysis with a flat reference and comparing the CNAs found with both the original and flat reference for the same samples.
How it works¶
CNVkit uses robust methods to extract a usable signal from the reference samples.
At each on– and off-target genomic bin, the read depths in each of the given normal samples are calculated and used to estimate the expected read depth and the reliability of this estimate. Specifically, CNVkit calculates Tukey’s biweight location, a weighted average of the normalized log2 coverages in each of the input samples, and biweight midvariance, the spread or statistical dispersion of read depth values using a similar weighting scheme. For background on these statistical methods see Lax (1985) and Randal (2008).
To adjust for the lower statistical reliability of a smaller number of samples for estimating parameters, a “pseudocount” equivalent to one sample of neutral copy number is included in the dataset when calculating these values.
If a FASTA file of the reference genome is given, for each genomic bin the fraction of GC (proportion of “G” and “C” characters among all “A”, “T”, “G” and “C” characters in the subsequence, ignoring “N” and any other ambiguous characters) and repeat-masked values (proportion of lowercased non-“N” characters in the sequence) are calculated and stored in the output reference .cnn file. For efficiency, the samtools FASTA index file (.fai) is used to locate the binned sequence regions in the FASTA file.
The same read-depth bias corrections used in the fix command are performed on each of the normal samples here. The result is a reference copy-number profile that can then be used to correct other individual samples.
Note
As with BAM files, CNVkit will automatically index the FASTA file if the
corresponding .fai file is missing or out of date. If you have copied the
FASTA file and its index together over a network, you may need to use the
touch command to update the .fai file’s timestamp so that CNVkit will
recognize it as up-to-date.
fix¶
Combine the uncorrected target and antitarget coverage tables (.cnn) and correct for biases in regional coverage and GC content, according to the given reference. Output a table of copy number ratios (.cnr).
cnvkit.py fix Sample.targetcoverage.cnn Sample.antitargetcoverage.cnn Reference.cnn -o Sample.cnr
How it works¶
The “observed” on- and off-target read depths are each median-centered and bias-corrected, as when constructing the reference. The corresponding “expected” normalized log2 read-depth values from the reference are then subtracted for each set of bins.
CNVkit filters out bins failing certain predefined criteria: those where the reference log2 read depth is below a threshold (default -5), the spread of read depths among all normal samples in the reference is above a threshold (default 1.0), or the RepeatMasker-covered proportion of the bin is above a threshold (default 99%).
A weight is assigned to each remaining bin depending on:
- The size of the bin;
- The deviation of the bin’s log2 value in the reference from 0;
- The “spread” of the bin in the reference.
(The latter two only apply if at least one normal/control sample was used to build the reference.)
Finally, the corrected on- and off-target bin-level copy ratios with associated weights are concatenated, sorted, and written to a .cnr file.
segment¶
Infer discrete copy number segments from the given coverage table:
cnvkit.py segment Sample.cnr -o Sample.cns
By default this uses the circular binary segmentation algorithm (CBS), which
performed best in our benchmarking. But with the -m option, the faster
HaarSeg
(haar) or Fused Lasso
(flasso) algorithms can be used instead.
If you do not have R or the R package dependencies installed, but otherwise do
have CNVkit properly installed, then haar will work for you. The other two
methods use R internally.
Fused Lasso additionally performs significance testing to distinguish CNAs from regions of neutral copy number, whereas CBS and HaarSeg by themselves only identify the supported segmentation breakpoints.
rescale¶
If there is a known level of normal-cell DNA contamination in the analyzed tumor sample (see the page on tumor heterogeneity), you can opt to rescale the log2 copy ratio estimates in your .cnr or .cns file to remove the impact of this contamination, so the resulting log2 ratio values in the file match what would be observed in a completely pure tumor sample.
The calculation of new log2 values for the sex chromosomes depends on the chromosomal gender of the sample and whether a male reference was used, while for autosomes the specified ploidy (default 2, diploid) is used. For example, with tumor purity of 60% and a male reference, letting CNVkit guess the sample’s chromosomal gender:
cnvkit.py rescale Sample.cns --purity 0.6 -y -o Sample.rescaled.cns
This can be done before or after segmentation, using a .cnr or .cns file; the resulting .cns file should be essentially the same.
The rescale command can also optionally re-center the log2 values, though
this will typically not be needed since the .cnr files are automatically
median-centered by the fix command when normalizing to a reference and
correcting biases. However, if the analyzed genome is highly aneuploid and
contains widespread copy number losses or gains unequally, median centering may
place copy-number-neutral regions slightly off-center from the expected log2
value of 0.0. To address such cases, alternative centering approaches can be
specified with the --center option:
cnvkit.py rescale Sample.cns --center mode
call¶
Given segmented log2 ratio estimates (.cns), round the copy ratio estimates to integer values using either:
- A list of threshold log2 values for each copy number state, or
- Rescaling for a given known tumor cell fraction and normal ploidy, then simple rounding to the nearest integer copy number.
cnvkit.py call Sample.cns -o Sample.call.cns
cnvkit.py call Sample.cns -y -m threshold -t=-1.1,-0.4,0.3,0.7 -o Sample.call.cns
cnvkit.py call Sample.cns -y -m clonal --purity 0.65 -o Sample.call.cns
The output is another .cns file, where the values in the log2 column are still log2-transformed and relative to the reference ploidy (by default: diploid autosomes, haploid Y or X/Y depending on reference gender). The segment log2 values are simply rounded to what they would be if the estimated copy number were an integer – e.g. a neutral diploid state is represented as 0.0, and a copy number of 3 on a diploid chromosome is represented as 0.58. The output .cns file is still compatible with the other CNVkit commands that accept .cns files, and can be plotted the same way with the scatter, heatmap and diagram commands.
To get the absolute integer copy number values in a human-readable form, use the
command export bed.
Calling methods¶
The “clonal” method uses the same calculation as the rescale command. It
considers the observed log2 ratios in the input .cns file as a mix of some
fraction of tumor cells (specified by --purity), possibly with altered copy
number, and a remainder of normal cells with neutral copy number (specified by
--ploidy for autosomes). This equation is rearranged to find the absolute
copy number of the tumor cells alone, rounded to the nearest integer. The
expected and observed ploidy of the sex chromosomes (X and Y) is different, so
it’s important to specify -y/--male-reference if a male reference was
used; the sample gender can be specified if known, otherwise it will be guessed
from the average log2 ratio of chromosome X.
The “threshold” method simply applies fixed log2 ratio cutoff values for each
integer copy number state. This method therefore does not require the tumor
cell fraction or purity to be known. The default cutoffs are reasonable for a
tumor sample with purity of at least 40% or so. For germline samples, the
-t values shown above may yield more accurate calls.
The thresholds work like:
| If \(\log_2\) value \(\leq\) | Copy number |
| -1.1 | 0 |
| -0.4 | 1 |
| 0.3 | 2 |
| 0.7 | 3 |
| ... | ... |
For homogeneous samples of known ploidy, you can calculate cutoffs from scatch by log-transforming the integer copy number values of interest, plus .5 (for rounding), divided by the ploidy. For a diploid genome:
>>> import numpy as np
>>> copy_nums = np.arange(5)
>>> print(np.log2((copy_nums+.5) / 2)
[-2. -0.4150375 0.32192809 0.80735492 1.169925 ]
Or, in R:
> log2( (0:4 + .5) / 2)
[1] -2.0000000 -0.4150375 0.3219281 0.8073549 1.1699250